Posted: November 14, 2016, 2:46pm
Last week I spent one of the most scientifically enriching weeks of my life at the Moffitt Cancer Center in Tampa Florida. The Integrated Math Oncology department (find them on Twitter here) hosted a workshop on resistance in cancer. This is a huge research focus in cancer and a very important problem to tackle. Moffitt’s point of view (which I whole-heartedly agree) is that these tough biological problems are best suited for teams of experimental biologists, practicing clinicians, and last but not least, math modelers like myself.

Our fearless leader of this workshop, Dr. Anderson, explains the need for an integrated approach to tackle the most difficult problems in cancer research. Each of five teams (red, blue, green, yellow, orange) had three team leaders from Moffitt: a biologist, clinician, and modeler. The teams would be supplemented by around 10 outside members from all over the world, and would spend the week creating a model of various cancers from the data provided by the team leaders. Our team (#TeamGreen) focused on skin cancer resistance to a drug called Pramlintide. We were given some microRNA and genetic expression data as a starting point and we were off to the races on the first day.
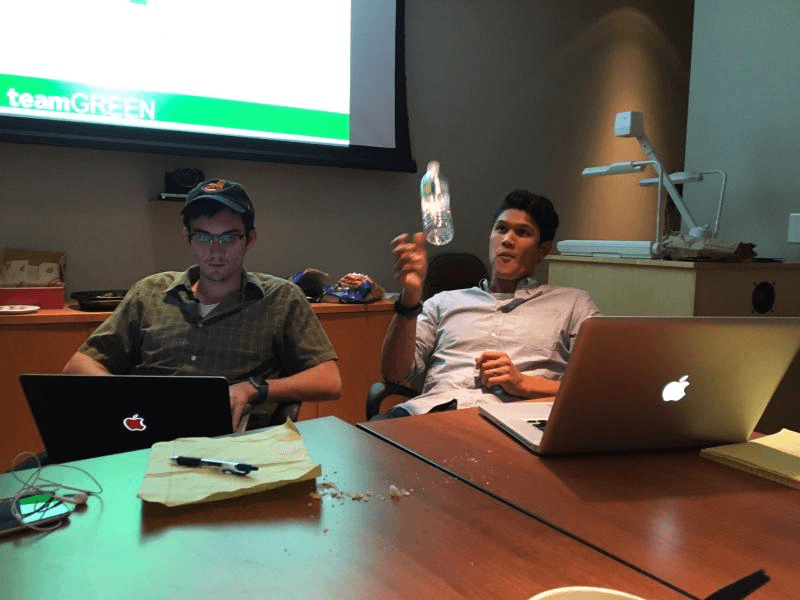

It had the feel of a hack-a-thon. We had a few team members well-trained in computer science and we largely spent the week coding. But the workshop was definitely more specialized than a typical code-a-thon because we had the valuable input of biologists interpreting data and informing the design of our models hand in hand all week long. It makes me wonder why science isn’t done in this format all the time! Work a 60 hour week one week in a strong team and then take the next week off? Sign me up!

Our (ambitious) project was designed with the intention of stratifying (separating) patients into different subgroups: who will be resistant to this promising new drug Pramlintide? Secondarily, a model may be useful in uncovering the dynamics and mechanisms of resistance. We decided on a neural network approach to go from microRNA and genetic influences on the phenotype of the cancerous cells. The drug targets metabolic pathways important in glycolytic activity.
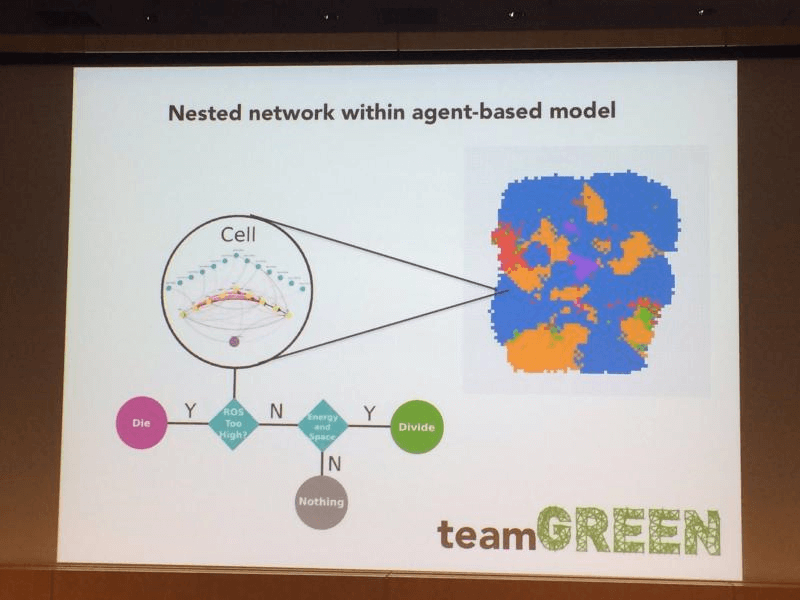
We plugged that neural network into each cell in an agent-based cellular automata model. This allows the cells to compete for resources and space on a grid lattice.
Overall I’m extremely thankful for the experience and the opportunity to participate in the competition. It gave me confidence to realize that I can contribute to a team of such remarkable and brilliant individuals. It made me admire Moffitt Cancer Center for their strong collaborations between varied disciplines toward a common goal. I would love to continue working with them in the future, should the opportunity arise!
Published on July 26th, 2017
Last updated on August 10th, 2017

